The purpose of this post is to demonstrate a first approach to the analysis of multiwavelength kinetic data, like those obtained using stopped-flow data. To practice, we will use data that were acquired during the stopped flow practicals of the
MetBio summer school from the
FrenchBIC. During the practicals, the student monitored the reaction of myoglobin (in its Fe(III) state) with azide, which yields a fast and strong change in the absorbance spectrum of the protein, which was monitored using a diode array. The data is publicly available on
zenodo.
Aims of this tutorial
The purpose of this tutorial is to teach you to use the free software
QSoas to run a simple, multiwavelength exponential fit on the data, and to look at the results. This is not a kinetics lecture, so that it will not go in depth about the use of the exponential fit and its meaning.
Getting started: loading the file
First, make sure you have a working version of QSoas, you can download them (for free)
there. Then download the data files from
zenodo. We will work only on the data file
Azide-1.25mm_001.dat, but of course, the purpose of this tutorial is to enable you to work on all of them. The data files contain the time evolution of the absorbance for all wavelengths, in a
matrix
format, in which each row correpond to a time point and each column to a wavelength.
Start
QSoas, and launch the command:
QSoas> load /comments='"'
Then, choose the
Azide-1.25mm_001.dat data file. This should bring up a horizontal red line at the bottom of the data display, with X values between about 0 and 2.5. If you zoom on the red line with the mouse wheel, you'll realize it is data. The
/comments='"' part is very important since it allows the extraction of the wavelength from the data. We will look at what it means another day. At this stage, you can look at the loaded data using the command:
QSoas> edit
You should have a window looking like this:
The rows each correspond to a data point displayed on the window below. The first column correspond to the X values, the second the Y values, and all the other ones to extra Y columns (they are not displayed by default). What is especially interesting is the first row, which contains a
nan as the X value and what is obviously the wavelength for all the Y values. To tell that QSoas should take this line as the wavelength (which will be the
perpendicular coordinate
, the coordinate of the other direction of the matrix), first close the edit window and run:
QSoas> set-perp /from-row=0
Splitting and fitting
Now, we have a single dataset containing a lot of Y columns. We want to fit all of them simultaneously with a (mono) exponential fit. For that, we first need to split the big matrix into a series of X,Y datasets (because fitting only works on the first Y). This is possible by running:
QSoas> expand /style=red-to-blue /flags=kinetics
Your screen should now look like this:
You're looking at the kinetics at all wavelengths at the same time (this may take some time to display on your computer, it is after all a rather large number of data points). The
/style=red-to-blue is not strictly necessary, but it gives the red to blue color gradient which makes things easier to look at (and cooler !). The
/flags=kinetics is there to attach a label (a
flag
) to the newly created datasets so we can easily manipulate all of them at the same time. Then it's time to fit, with the following command:
QSoas> mfit-exponential-decay flagged:kinetics
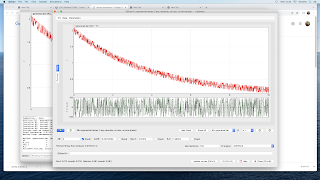
This should bring up a new window. After resizing it, you should have something that looks like this:
The bottom of the fit window is taken by the parameters, each with two checkboxes on the right to set them fixed (i.e. not determined by the fitting mechanism) and/or global (i.e. with a single value for all the datasets, here all the wavelengths). The top shows the current dataset along with the corresponding fit (in green), and, below, the residuals. You can change the dataset by clicking on the horizontal arrows or using Ctrl+PgUp or Ctrl+PgDown (keep holding it to scan fast). See the
Z = 728.15 showing that QSoas has recognized that the currently displayed dataset corresponds to the wavelength 728.15. The equation fitted to the data is: $$y(x) = A_\infty + A_1 \times \exp -(x - x_0)/\tau_1$$
In this case, while the \(A_1\) and \(A_\infty\) parameters clearly depend on the wavelength, the time constant of evolution should be independent of wavelength (the process happens at a certain rate regardless of the wavelength we're analyzing), so that the \(\tau_1\) parameter should be common for all the datasets/wavelengths. Just click on the
global
checkbox at the right of the
tau_1 parameter, make sure it is checked, and hit the
Fit
button...
The fit should not take long (less than a minute), and then you end up with the results of the fits: all the parameters. The best way to look at the non global parameters like \(A_1\) and \(A_\infty\) is to use the
Show Parameters
item from the
Parameters
menu. Using it and clicking on
A_inf too should give you a display like this one:
The
A_inf parameter corresponds to the spectum at infinite time (of azide-bound heme), while the
A_1 parameter corresponds to the difference spectrum between the initial (azide-free) and final (azide-bound) states.
Now, the fit is finished, you can save the parameters if you want to reload them in a later fit by using the
Parameters
/
Save
menu item or export them in a form more suitable for plotting using
Parameters
/
Export
(although QSoas can also display and the parameters saved using
Save
). This concludes this first approach to fitting the data. What you can do is
How to read the code above
All the lines starting by
QSoas> in the code areas above are meant to be typed into the QSoas command line (at the bottom of the window), and started by pressing
enter at the end. You must remove the
QSoas> bit. The other lines (when applicable) show you the response of QSoas, in the terminal just above the command-line. You may want to play with the
QSoas tutorial to learn more about how to interact with QSoas.
About QSoas
QSoas is a powerful open source data analysis program that focuses on flexibility and powerful fitting capacities. It is released under the
GNU General Public License. It is described in
Fourmond, Anal. Chem., 2016, 88 (10), pp 5050–5052. Current version is
3.1. You can freely (and at no cost) download its source code or precompiled versions for MacOS and Windows
there. Alternatively, you can clone from the
GitHub repository.
Contact: find my email address
there, or contact me on
LinkedIn.